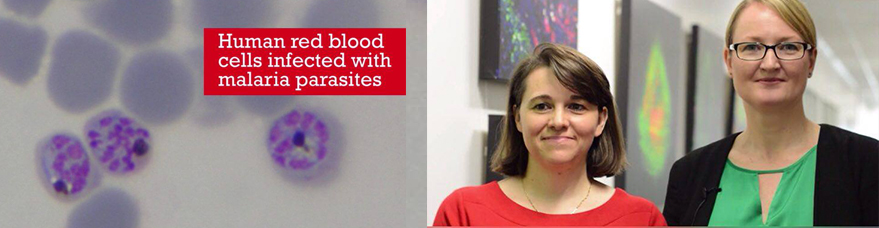
Malaria remains one of the world’s leading causes of mortality in developing countries. Last year alone, it killed more than 400,000 people, mostly young children. This week in ACS Central Science, an international consortium of researchers unveils the mechanics and findings of a unique “open science” project for malaria drug discovery that has been five years in the making.

The current gold standard antimalarial treatments are based on artemisinin, a compound developed in the 1970s in China, combined with a partner drug. Yet, resistance to artemisinin and its partners has already emerged in some parts of the world. If it spreads, there are no viable replacements. Given the lack of commercial incentive for industry to develop drugs for neglected diseases, and because academic researchers often lack resources to move compounds forward, there is a clear need for new approaches. In response, Matthew Todd from The University of Sydney together with the not-for-profit research and development organization Medicines for Malaria Venture proposed an “open source” solution akin to the open source concept used in software development.
The Eskitis Institute’s Professor Vicky Avery, who’s lab contributed to the biological profiling of compounds, said the work leads the way in a new paradigm in how we undertake drug discovery, especially for neglected diseases.
Professor Avery adds, “Throughout our involvement since 2011, it has been interesting to see how this approach which has been driven by Mat has evolved. He should be congratulated for his incredible perseverance and commitment to open source drug discovery.
“This is another example of where scientists from around the world have joined forces to enhance/further drug discovery for one of the world’s biggest killers.”
More than 50 researchers from 21 organizations in eight countries added their research to the project, which started with a large set of potential drug molecules made public by the company GlaxoSmithKline. Anyone willing to contribute — anywhere in the world — was welcome to share data and collaborate by adding comments to an electronic notebook as part of the Open Source Malaria Consortium. Some scientists designed and synthesized new generations of the antimalarial compounds; others ran assays and interpreted results. Several rounds of research were conducted, addressing water solubility and structural issues, with all the data being made public in real time. A wide array of scientists, from professors through to undergraduates, participated by choice, agreeing that no one would individually seek patents to protect their contributions. The authors note that the current results, while promising, are merely the beginning of the story. They continue to welcome additional contributions, also researched openly and collaboratively.
The authors acknowledge funding from Medicines for Malaria Venture, the Australian Research Council, the Australian National Health and Medical Research Council, the European Molecular Biology Laboratory member states, the National Science Foundation and the Wellcome Trust.
Original source: “Open Source Drug Discovery – Highly Potent Antimalarial Compounds Derived from the Tres Cantos Arylpyrroles”
Know more: Eskitis Institute for Drug Discovery
The Open Source Malaria Project